Tine Huyse, Researcher, Department of Biology
Royal Museum for Central Africa, Tervuren, Belgium
Human migration can lead to the introduction and mixing of culture, music and languages (Maniacky, 2009), but it can also lead to the introduction of new fauna and flora that can become established in the new area and interact with local species (Nova and Mack, 2001). It can however also lead to the introduction of diseases. Every human being is parasitized by several species of bacteria, viruses or helminths, often invisible to the naked eye. While contagious diseases can easily spread through direct human contact (e.g. measles), many infectious diseases depend on the presence of a vector to be transmitted. Examples include mosquito species that transmit Plasmodium parasites in case of malaria, or fresh water snails transmitting blood flukes in case of schistosomiasis (also known as snail fever). The latter will be the focus of this contribution.
It has been hypothesized that Schistosoma mansoni was introduced during the Trans-Atlantic slave trade that started in the 16th century.
Schistosomiasis or bilharzia is an important neglected tropical disease that affects more than 200 million people worldwide, of which at least 90% live in sub-Saharan Africa (WHO, 2014). Symptoms include abdominal pain, splenomegaly, haematuria and liver fibrosis and sometimes can be fatal. The disease is caused by parasitic worms of the genus Schistosoma. These flukes live in the blood veins of mammals where they feed on red blood cells. There they continuously produce eggs that either pass with the urine or faeces to continue the lifecycle, or they become trapped in the surrounding tissues, liver and spleen, where they cause local inflammation. The eggs will hatch in the water and infect a freshwater snail, in which it will develop into cercariae that are infective for man (Gryseels et al., 2006). Of the seven Schistosoma species that infect humans, Schistosoma mansoni has the widest distribution. It is present in 54 countries, mostly on the African continent but it is the only schistosome species that occurs on the South American continent (Figure 1). It has been hypothesized that it was introduced during the Trans-Atlantic slave trade that started in the 16th century. Due to the presence of a suitable snail host in South America, S. mansoni could successfully establish itself and spread across the continent.
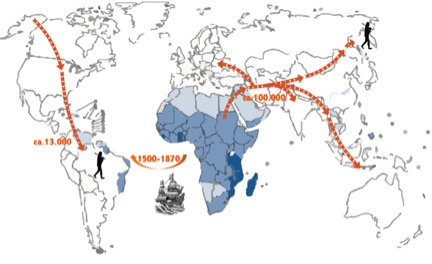
Figure 1. While modern humans had to cross the Bering Strait to reach South America it has been hypothesized that the parasite was introduced from West Africa during the Trans-Atlantic slave trade. Timings are approximate.
For as far as possible, this assumption is confirmed by archeo-parasitological studies. There have been no records of schistosome eggs in pre-Columbian archeological material in the New World, while eggs of the human nematode Trichuris trichiura have been found in several archeological sites in Brazil, Chile and Peru dating 8000 – 950 BP (Leles et al., 2008). In contrast, there are several records in the Old World of schistosome eggs in Egyptian or Sudanese mummies dating from several millennia B.C. (Gonçalves et al., 2003). Absence of paleoparasitological data might however not be conclusive evidence that schistosomiasis was not present before the slave trade commenced. The recent development of molecular genotyping techniques for helminth parasites provide an alternative tool to determine the provenance of South American schistosome strains and test the hypothesis of a recent introduction from West Africa.
The first study to generate molecular sequence data for a wide geographical range of S. mansoni isolates was performed by Morgan et al. (2005). They sequenced mitochondrial DNA fragments of schistosome populations collected in 17 countries. Subsequent phylogenetic analyses firmly clustered West African and South American schistosome populations. The fact that genetic distances between those populations was smaller than the distance between West African and East African populations clearly demonstrates a recent colonization from the New World by West African strains, ‘almost certainly with the African slave trade of the 15th to 19th century’ (Morgan et al., 2005). East African populations displayed the highest mitochondrial DNA diversity whereas the lowest diversity was found in the New World. This suggests that S. mansoni originated in East Africa, around 300-430.000 years ago according to the molecular clock analysis.
There were a few drawbacks or limitations in the study design. The first limitation is linked with the nature of the molecular marker that was used, namely mitochondrial DNA sequences. The mitochondrial DNA is haploid because it is only passed along the maternal line (uniparental marker), and therefore less informative than nuclear markers (biparental markers). The authors could not conclude whether the low diversity in South American schistosome populations was due to a demographic bottleneck or due to natural selection as they only included mitochondrial DNA sequence data. A demographic bottleneck would have occurred if only a reduced number of parasites survived transatlantic crossing. In that scenario only a small number of schistosome strains would seed the new population in South America resulting in a lower genetic diversity compared to Africa (Figure 2). Another possibility is that many parasite strains successfully crossed the ocean, but that only a few of them could adapt to the new snail host in South America (Figure 3). This new environment could pose a strong selection pressure resulting in a selective sweep where only a few strains can survive. The only way to discriminate amongst both scenarios is the additional use of polymorphic nuclear markers like microsatellite markers. These are short tandem repeat sequences in the nuclear genome (biparental) with a high mutation rate and therefore provide more resolution than mitochondrial DNA sequences (Morin et al., 2004).
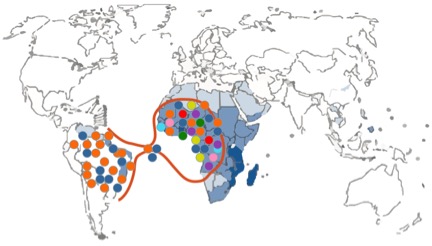
Figure 2. Schematic representation of a demographic bottleneck. Different colors depict different genetic schistosome strains.
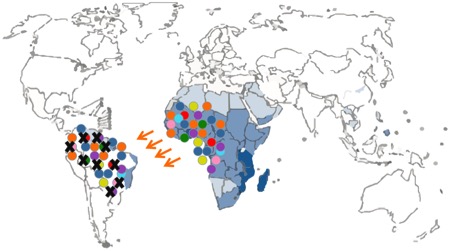
Figure 3. Schematic representation of a selective sweep. Different colors depict different genetic schistosome strains; a black cross represents extinction due to natural selection.
The second limitation of the study was linked with the biological samples. The molecular analyses were performed on lab-passaged material, i.e. adult schistosome worms obtained after experimental infection of laboratory mice by cercariae isolated from field-collected freshwater snails (Morgan et al., 2005). This methodology results in a decreased genetic diversity of the resulting parasite population due to selection and/or founder effects during experimental passage (Bech et al., 2010). Molecular analyses directly on field-collected cercarae or miracidia are therefore preferred. Another drawback is the sampling design, which is focused on East-Africa, while countries that were likely to be involved in the slave trade, like DR Congo, Angola, Gabon, Nigeria, Liberia or Ivory Coast are missing. Therefore new markers and natural S. mansoni isolates from additional geographic regions should be included in future studies.
With the development of new collection and extraction protocols of schistosome field isolates (Gower et al., 2007; Van den Broeck et al., 2011), microsatellite genotyping on individual specimens are now routinely performed (Van den Broeck et al., 2011; Gower et al., 2012). By comparing genetic diversity of S. mansoni populations from Senegal and Brazil both at the nuclear (microsatellites) and the mitochondrial level (cytochrome oxidase 1 partial sequences), we find evidence for the selective sweep hypothesis outlined above (personal data). We are currently increasing our sampling range within West Africa to obtain a higher resolution in order to trace the provenance of Brazilian schistosome populations.
Molecular epidemiological and linguistic studies could complement each other in the study on historical links between South America and Africa.
When parasites from disparate geographic localities are introduced in the same area, they can mix and reproduce, which can create new genotypes and possibly new phenotypes. This admixture between different source populations has been recently demonstrated for Plasmodium falciparum, the causative agent of malaria in humans. Using the same method and type of DNA markers they found evidence for two different introduction routes in South America, followed by admixture (Yalcindag et al., 2010). The authors linked this with the subdivision of the Americas during the 16th-19th century into the Spanish and the Portuguese empires. The Spanish ferried most slaves to the West Indies colonies (Jamaica, Cuba, and other islands), while the Portuguese shipped to Rio de Janeiro and Salvador de Bahia. It remains to be tested whether we find the same pattern for schistosomes.
As a concluding remark in order to imbed this study in the wider scope of the Africa beyond Africa conference, one could draw the parallel between the mixing of alleles and the mixing of languages, which could both lead to the creation of something new, be it phenotypes or words, still carrying the signature of their respective origin. Therefore molecular epidemiological and linguistic studies could complement each other in the study on historical links between South America and Africa.
References
Bech, N., Beltran, S., Portela, J., Rognon, A., Allienne J-F, Boissier, J. & Théron, A. 2010. Follow-up of the genetic diversity and snail infectivity of a Schistosoma mansoni strain from field to laboratory. Infect. Genet. Evol. 10: 1039–1045
Garrigan, D. & M.F. Hammer 2006. Reconstructing human origins in the genomic era. Nature Reviews Genetics 7: 669–680
Gonçalves M.L.C., Araújo A. & Ferreira L.F. 2003. Human intestinal parasites in the past: New findings and a review. Mem Inst Oswaldo Cruz 98: 103-118
Gower, C.M., Shrivastava, J., Lamberton, P.H., Rollinson, D., Webster, B.L., Emery, A., Kabatereine, N.B., & Webster, J.P., 2007. Development and application of an ethically and epidemiologically advantageous assay for the multi-locus microsatellite analysis of Schistosoma mansoni. Parasitology 134, 523–536.
Gower, C.M., Gouvras A.N., Lamberton, P.H. et al. 2013. Population genetic structure of Schistosoma mansoni and Schistosoma haematobium from across six sub-Saharan African countries: Implications for epidemiology, evolution and control. Acta Tropica, 128, 261-274
Gryseels, B., Polman, K., Clerinx, J. & Kestens, L. 2006. Human schistosomiasis. Lancet 368, 1106–1118
Leles D., Araújo A., Ferreira L.F., Vicente A.C.P., & Iñiguez A.M. 2008. Molecular paleoparasitological diagnosis of Ascaris sp. from coprolites: new scenery of ascariasis in pre-Columbian South America times. Mem Inst Oswaldo Cruz 103: 106-108
Maniacky, J. 2009. Thèmes régionaux bantu et africanismes brésiliens. In: Margarida Petter & Ronald Beline Mendes (eds), Proceedings of the Special World Congress of African Linguistics São Paulo 2008. Exploring the African Language Connection in the Americas. São Paulo: Humanitas, pp. 153-165. (PR) ISBN: 978-85-7732-121-6
Morin, P. A., Luikart, G., Wayne, R. K. & SNP workshop group. 2004. SNPs in ecology, evolution and conservation. TREE, 19, 208–216
Novak, S.J. & Mack, R.N. 2001. Tracing plant introduction and spread: genetic evidence from Bromus tectorum (Cheatgrass). Bioscience, 51, 114–122
Van den Broeck, F., Geldof, S., Polman, K., Volckaert, F.A.M., & Huyse, T., 2011. Optimal sample storage and extraction procotols for reliable multilocus genotyping of the human parasite Schistosoma mansoni. Infect. Genet. Evol. 11, 1413–1418.
WHO (2014) Schistosomiasis fact sheet number 115. http://www.who.int/mediacentre/factsheets/fs115/en/index.html.
Yalcindag E., Elguero E., Arnathau C. et al. 2012. Multiple inde-pendent introductions of Plasmodium falciparum in South America. PNAS, 109, 511–516